The problem with healthcare analytics
Healthcare organizations sit on valuable data, but sharing it is a minefield. A hospital wants to study acute kidney injury rates across its ICU population. An external research team has the statistical expertise. The data exists — hundreds of thousands of lab observations stored as FHIR resources in Aidbox. But handing over raw patient records violates HIPAA.
The usual workaround is a bespoke ETL pipeline: export data, run a script to strip identifiers, hope nothing slips through, load the scrubbed data somewhere else. This is slow, error-prone, and hard to audit. Every new research question means another pipeline.
What if the de-identification happened inside the database, declared alongside the view that shapes the data?
De-identification as a ViewDefinition feature
Aidbox now supports per-column de-identification in ViewDefinitions. You annotate columns with a FHIR extension that specifies which transformation to apply. The SQL compiler wraps each column expression with a PostgreSQL function — sensitive data is transformed before it ever reaches the output.
{
"resourceType": "ViewDefinition",
"name": "deident_observations",
"status": "active",
"resource": "Observation",
"select": [{
"column": [
{
"name": "patient_id",
"path": "subject.getReferenceKey(Patient)",
"extension": [{
"url": "http://health-samurai.io/fhir/core/StructureDefinition/de-identification",
"extension": [
{"url": "method", "valueCode": "cryptoHash"},
{"url": "cryptoHashKey", "valueString": "my-secret-key"}
]
}]
},
{
"name": "status",
"path": "status"
}
]
}]
}
The patient_id column carries a de-identification extension. It will output an HMAC-SHA256 hash instead of the real patient reference — deterministic (same patient always gets the same hash) and irreversible (you cannot recover the original value from the hash). The status column has no extension and passes through unchanged.
The ViewDefinition Builder in Aidbox UI makes this visual — click the shield icon on any column to pick a method and set parameters:
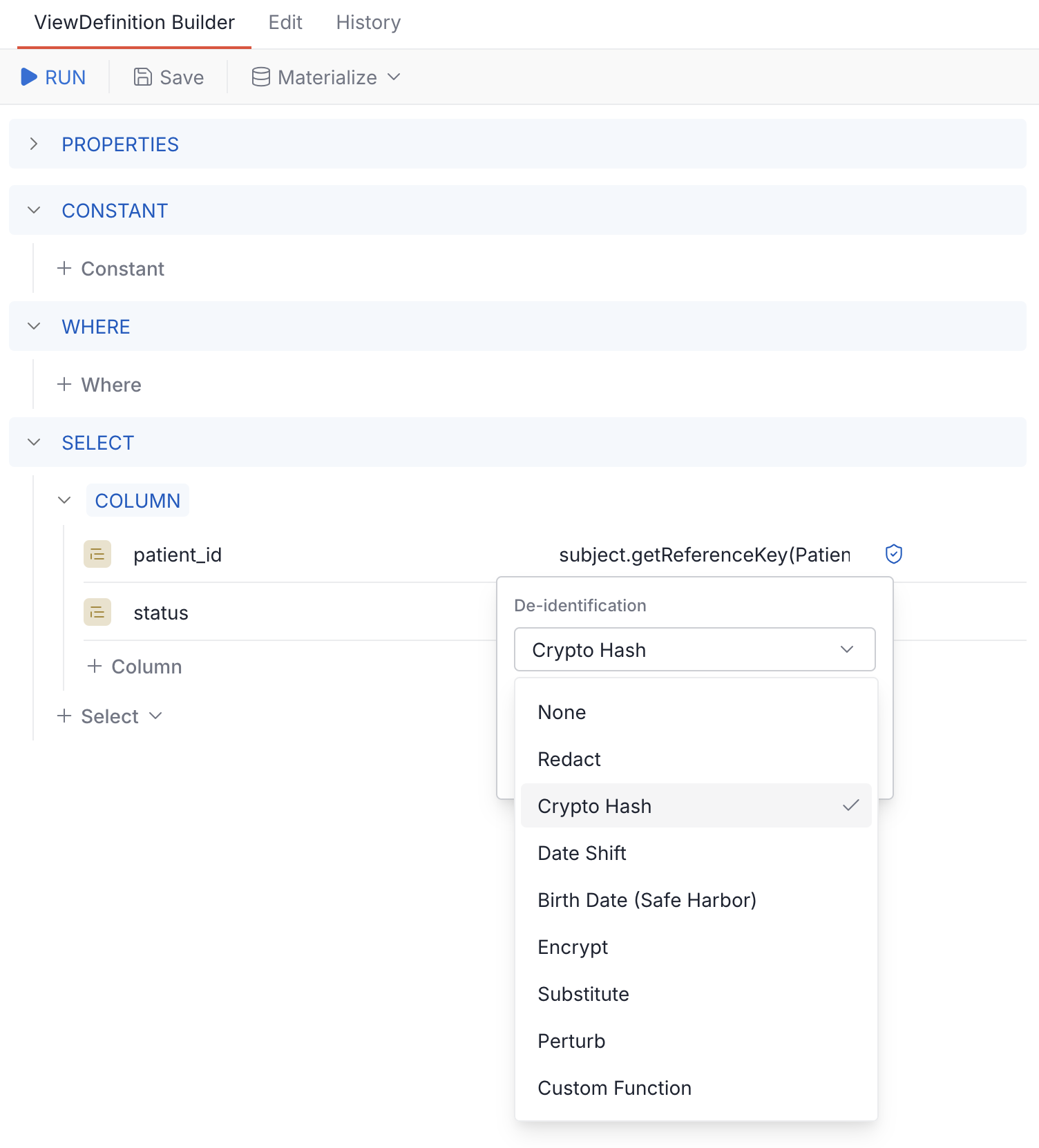
Available methods
| Method | What it does | Key parameters |
|---|---|---|
| redact | Replace with NULL | — |
| cryptoHash | HMAC-SHA256 hash (deterministic, one-way) | cryptoHashKey |
| dateshift | Shift dates by ±1–50 days per resource | dateShiftKey |
| birthDateSafeHarbor | Date shift + auto-redact when age >89 (Patient.birthDate only) | dateShiftKey |
| encrypt | AES-128-CBC, base64 output (reversible with key) | encryptKey |
| substitute | Replace with a fixed string | replaceWith |
| perturb | Add random noise to numeric values | span, rangeType, roundTo |
| custom_function | Call your own PostgreSQL function | custom_function, custom_arg (optional) |
Two methods are especially relevant for the pattern we're about to explore:
cryptoHash produces the same output for the same input — which means you can use it to join tables. Hash the patient ID in two different ViewDefinitions with the same key, and the hashes will match. This is the foundation for re-identification.
dateshift shifts all dates within the same resource by the same offset, preserving temporal relationships. A 3-day gap between two events stays a 3-day gap. The offset is derived from HMAC(dateShiftKey, resource.id), so it's deterministic per resource but unpredictable without the key.
Practical example: outsourcing AKI research
Let's walk through a realistic scenario using MIMIC-IV data loaded into Aidbox — 100 ICU patients, over 800,000 observations.
The goal: A hospital wants an external research team to analyze acute kidney injury (AKI) rates in its ICU population. The researchers need creatinine lab results but must not see real patient identifiers or exact dates.
The approach: Create two materialized tables — one de-identified dataset for the researchers, and one internal mapping table that connects hashed IDs back to real patients.
Designing the de-identified view
We start with a ViewDefinition on the Observation resource, filtered to creatinine lab results. Four columns, each with a different de-identification strategy:
Patient ID — the column most in need of protection. We apply cryptoHash to turn the real UUID into an irreversible HMAC-SHA256 hex string:
// ---8<--- snip
{
"name": "patient_id",
"path": "subject.getReferenceKey(Patient)",
"extension": [{
"url": "..de-identification",
"extension": [
{"url": "method", "valueCode": "cryptoHash"},
{"url": "cryptoHashKey", "valueString": "aki-study-2026"}
]
}]
}
// ---8<--- snap
Creatinine value and unit — the clinical data the researcher actually needs. No de-identification — lab values are not identifiers:
// ---8<--- snip
{
"name": "creatinine",
"path": "value.ofType(Quantity).value",
"type": "decimal"
},
{
"name": "unit",
"path": "value.ofType(Quantity).unit"
}
// ---8<--- snap
Effective date — shifted by a deterministic per-resource offset using dateshift. A 3-day gap between two labs stays a 3-day gap, but the real calendar date is hidden:
// ---8<--- snip
{
"name": "effective_date",
"path": "effective.ofType(dateTime)",
"extension": [{
"url": "..de-identification",
"extension": [
{"url": "method", "valueCode": "dateshift"},
{"url": "dateShiftKey", "valueString": "aki-study-2026"}
]
}]
}
// ---8<--- snap
The mapping table
The second ViewDefinition is for re-identification — it stays internal. Two columns on the Patient resource, both pointing to id: one plain, one hashed with the same key as above. Because cryptoHash is deterministic, the hashed IDs will match across both tables.
Building in the UI
The ViewDefinition builder makes this visual. Click the shield icon on any column to pick a de-identification method and set its parameters:
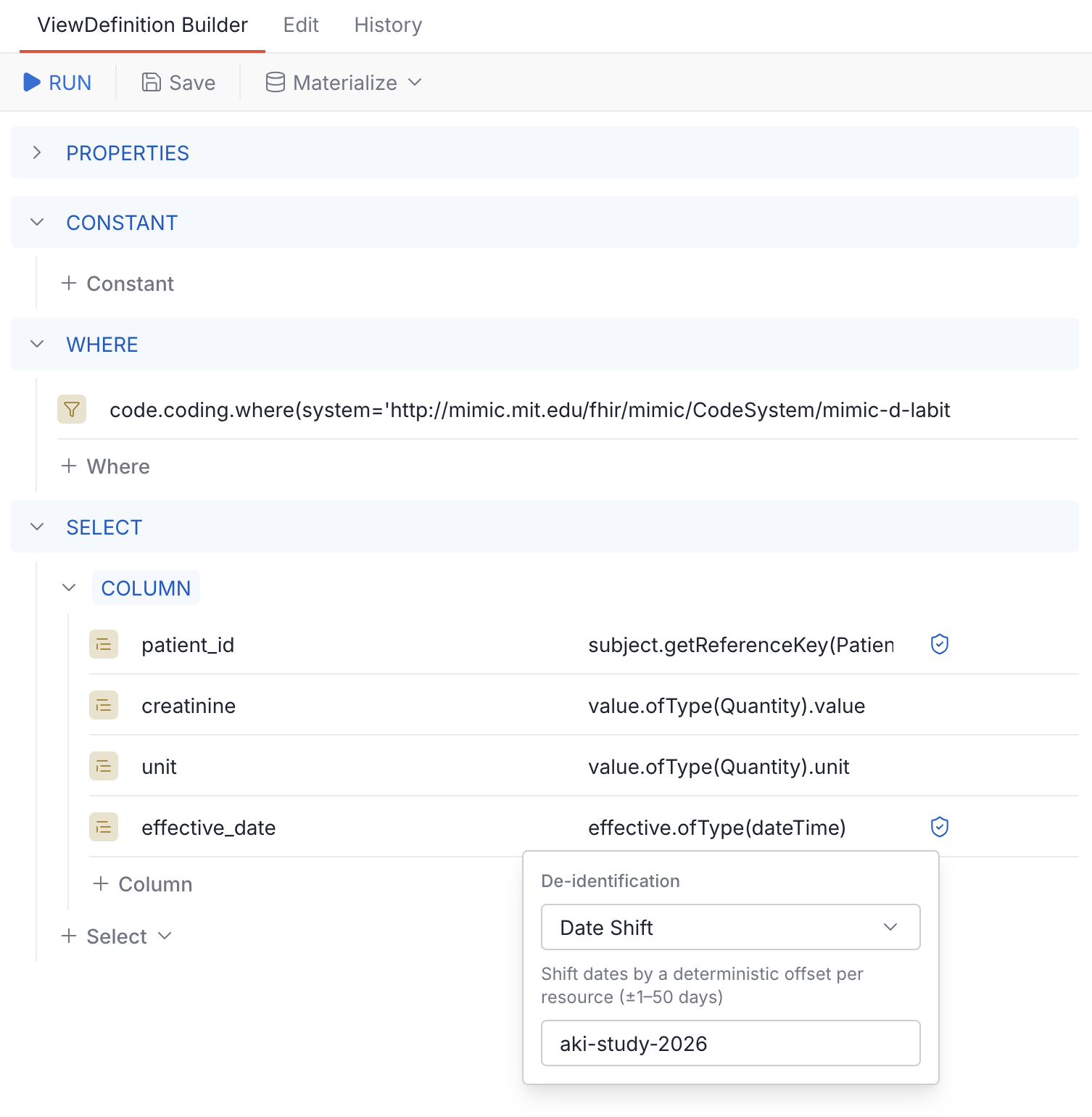
Once saved, materialize as a table:
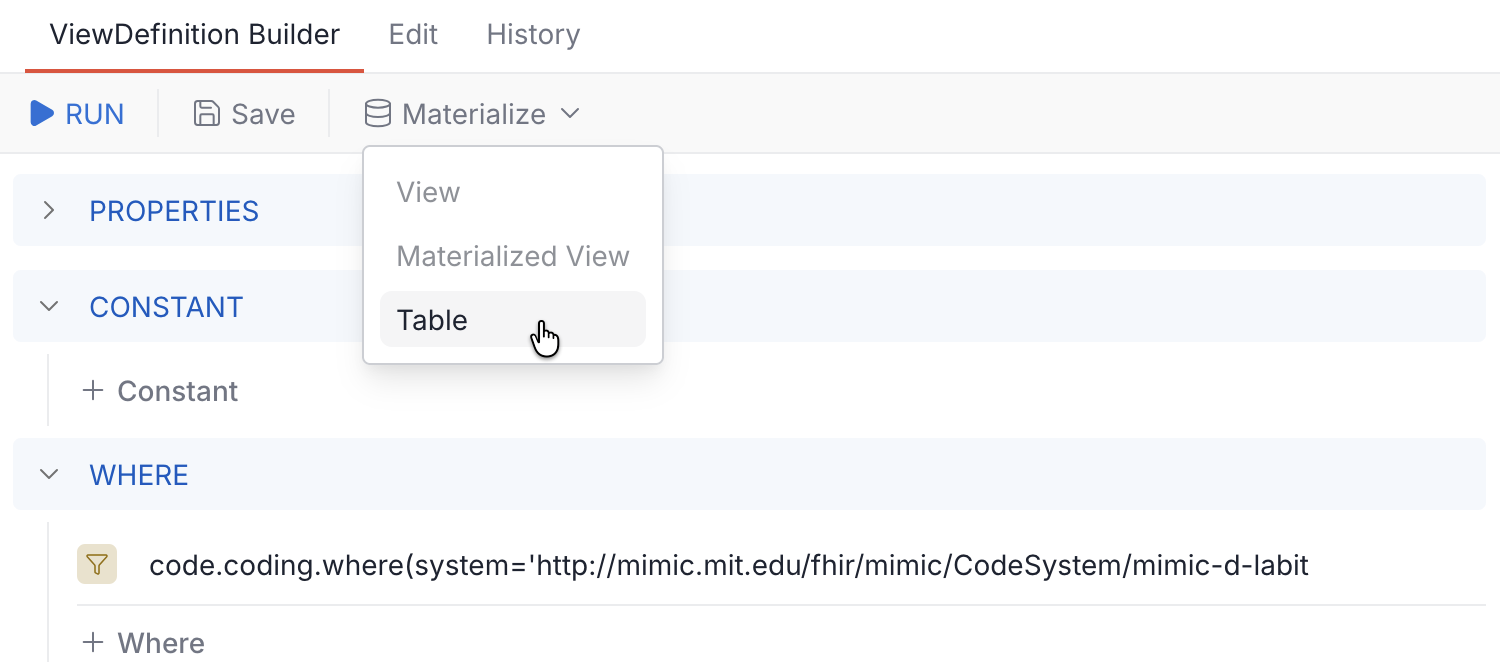
ViewDefinitions with de-identification extensions can only be materialized as table — not view or materialized-view. PostgreSQL stores the full SQL definition of views in system catalogs (pg_views), which would expose your cryptographic keys to anyone with catalog access. Tables store only the transformed data.
What the researcher sees
The researcher queries the de-identified table. No real patient IDs, no real dates:
SELECT * FROM sof.deident_creatinine LIMIT 5;
patient_id | creatinine | unit | effective_date |
|---|---|---|---|
| db1d98d58...f1508c7b439aba | 1.1 | mg/dL | 2113-01-17T09:45:00-05:00 |
| db1d98d58...f1508c7b439aba | 1.4 | mg/dL | 2116-03-05T16:16:00-05:00 |
| f852c0e57...ca9c6e3db462a9 | 0.8 | mg/dL | 2153-03-08T02:17:00-04:00 |
| 59e60d080...01688175f47b16 | 1.7 | mg/dL | 2147-09-09T15:20:00-04:00 |
| 6283390bd...738cd02dac4c08 | 0.8 | mg/dL | 2140-10-03T15:20:00-04:00 |
The patient_id is a hex string — the researcher can group by it, count distinct patients, track trends over time. But they cannot reverse it to find the real patient.
Computing the metric
AKI is commonly identified by elevated creatinine. A simplified screening criterion: creatinine above 1.5 mg/dL indicates possible kidney injury. The researcher runs:
SELECT
CASE
WHEN creatinine <= 1.1 THEN 'Normal (≤1.1)'
WHEN creatinine <= 1.5 THEN 'Elevated (1.1–1.5)'
WHEN creatinine <= 3.0 THEN 'High (1.5–3.0)'
ELSE 'Critical (>3.0)'
END AS risk_category,
count(*) AS observation_count,
count(DISTINCT patient_id) AS patient_count,
round(avg(creatinine)::numeric, 2) AS avg_creatinine
FROM sof.deident_creatinine
GROUP BY 1
ORDER BY 1;
risk_category | observation_count | patient_count | avg_creatinine |
|---|---|---|---|
| Critical (>3.0) | 307 | 19 | 5.08 |
| Elevated (1.1–1.5) | 393 | 40 | 1.33 |
| High (1.5–3.0) | 575 | 30 | 2.11 |
| Normal (≤1.1) | 1728 | 84 | 0.77 |
19 patients had creatinine readings above 3.0 mg/dL — a critical threshold suggesting severe kidney injury. The researcher identifies these patients by their hashed IDs and sends the results back to the hospital.
Re-identifying for clinical follow-up
The hospital joins the research results against the internal mapping table — matching hashed IDs to real patient UUIDs:
SELECT
m.real_id,
max(o.creatinine) AS peak_creatinine,
count(*) AS high_readings
FROM sof.deident_creatinine o
JOIN sof.patient_id_map m ON o.patient_id = m.hashed_id
WHERE o.creatinine > 3.0
GROUP BY m.real_id
ORDER BY peak_creatinine DESC
LIMIT 20;
real_id | peak_creatinine | high_readings |
|---|---|---|
| af0e5009-d87d-52a8-ac8a-676e471c41f1 | 15.2 | 9 |
| 8e77dd0b-932d-5790-9ba6-5c6df8434457 | 11 | 80 |
| 1cf9e585-806c-513b-80af-4ca565a28231 | 9.1 | 62 |
| 568cb149-804c-59e8-bdf5-816e8151cd22 | 8.3 | 9 |
| df756e08-6ea8-5d69-b918-67911945f827 | 7.9 | 11 |
| 7ec7078a-0593-5a99-9862-ebbff47fd1c5 | 6.4 | 20 |
| dd2bf984-33c3-5874-8f68-84113327877e | 6.2 | 26 |
| 4f773083-7f4d-5378-b839-c24ca1e15434 | 5.3 | 11 |
| 72d56b49-a7ee-5b9a-a679-25d1c836d3c3 | 5.1 | 3 |
| ... | ... | ... |
| 842680b3-e421-58cc-8050-3b29668b438c | 3.2 | 1 |
Now the hospital knows exactly which patients need clinical attention — without ever having exposed their identities to the research team.
The full picture
The key insight: the same cryptoHashKey ties everything together. The researcher works entirely with hashed identifiers. The hospital holds the mapping table and the key. Re-identification happens only internally, only when needed.
Pre-built Safe Harbor ViewDefinitions
Writing de-identification rules for every resource type is tedious. We provide io.health-samurai.de-identification.r4 — a FHIR package available through Aidbox's artifact registry with ready-made ViewDefinitions for 17 common R4 resource types: Patient, Encounter, Observation, Condition, Claim, ExplanationOfBenefit, AllergyIntolerance, DiagnosticReport, MedicationRequest, MedicationDispense, MedicationAdministration, Immunization, Procedure, Specimen, DocumentReference, Practitioner, and Location.
Each ViewDefinition applies Safe Harbor rules:
- cryptoHash on identifiers and references — consistent across tables for joins
- dateshift on clinical dates — preserves temporal relationships
- birthDateSafeHarbor on Patient.birthDate — date shift plus automatic redaction when the patient is over 89, as required by 45 CFR 164.514(b)(2)(i)(C)
- redact on names, addresses, and other direct identifiers
All cryptographic key parameters are blank by default — set your own keys before materializing. Install the package via FAR, configure keys, materialize as tables, and start querying.
Getting started
- Enable fhir-schema mode — set
fhir.validation.fhir-schema-validation=true(required since Aidbox 2604) - Install
io.health-samurai.de-identification.r4via FAR, or write your own ViewDefinitions - Set cryptographic keys in each ViewDefinition — blank keys will be rejected
- Materialize as tables —
$materializewithtype=tableor via the ViewDefinition builder - Query with SQL from Aidbox SQL console, Grafana, or any BI tool
For full method reference and parameter details, see the de-identification documentation. To try the complete scenario from this post with a single docker compose up, check out the de-identification example.






